Labrador
| Function | A web based tool to manage and automate the processing of publicly available datasets. |
|---|---|
| Language | PHP, MySQL, Javascript |
| Requirements |
A suitable web server with PHP installed (apache config files are included) A MySQL server |
| Code Maturity | Beta - Being used in production, but still under active development. |
| Code Released | Yes, under GPL v3 or later. |
| Initial Contact | Phil Ewels |
Download Now |
|
End-User Usage Tutorial
Screencast showing basic usage of Labrador
Administration Tutorial
Screencast showing administrative use of Labrador
Installation Walkthrough
Screencast of installing Apache, PHP, MySQL and Labrador on a blank server
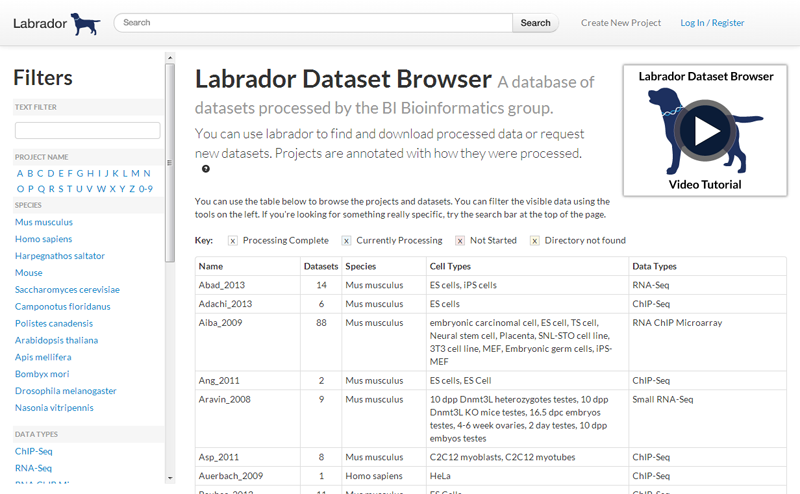
Labrador is a web based tool to manage projects and automate the processing of publicly available datasets.
Researchers can use Labrador to:
- Browse and search previously processed datasets
- View processing and analysis reports in their web browser
- Download data through their web browser
- Request new datasets, with required information automatically retrieved from accession numbers
Bioinformaticians can use Labrador to:
- Speed up retrieval of project information from repositories
- Catalogue processing and analysis
- Create automated analysis bash scripts
- Customise templates for analysis script generation
Project metadata is stored within a mysql database. Code is provided with some example analysis script pipelines which can be easily modified to work with existing workflows. This analysis script generation can help to standardise in-house processing and streamline pipelines.
Documentation
The Labrador documentation is available online and comes bundled with Labrador.
Changelog
- 26-04-10: Version 0.1 released
-
- Initial Labrador release. Everything works on our test systems, though more development is expected.