Compter
| Function | Compter is a program which quantitates and visualises compositional biases in sets of DNA or RNA sequences. |
|---|---|
| Language | Perl and R |
| Requirements | The Pheatmap R package. A web server if you want to use the web interface (optional) |
| Code Maturity | Beta - still being developed but is fully functional in its current form |
| Code Released | Yes, under GPL v3 or later. |
| Initial Contact | Simon Andrews |
Download Now |
|

Understanding the composition of a set of sequences can be an important step in the analysis of many different kinds of data. The underlying cause of the selection of a set of genomic locations can often be influenced by sequence composition and this could be the result of either real biology or technical artefacts
If you have a single set of sequences it can be useful to know if it is enriched for certain sequences compared to the genome as a whole. It can also be useful to know if your set of sequences all behaves similarly or if there are sub-groups within it with different compositional biases.
If you have multiple sets of sequences you may wish to see if they have consistent compositional differences between them before getting into more complicated functional analyses such as motif searching.
A compositional analysis can be a useful part of many different analysis pipelines including RNA-Seq, ChIP-Seq methylation analysis and many other situations which generate a list of candidate genes or genomic regions.
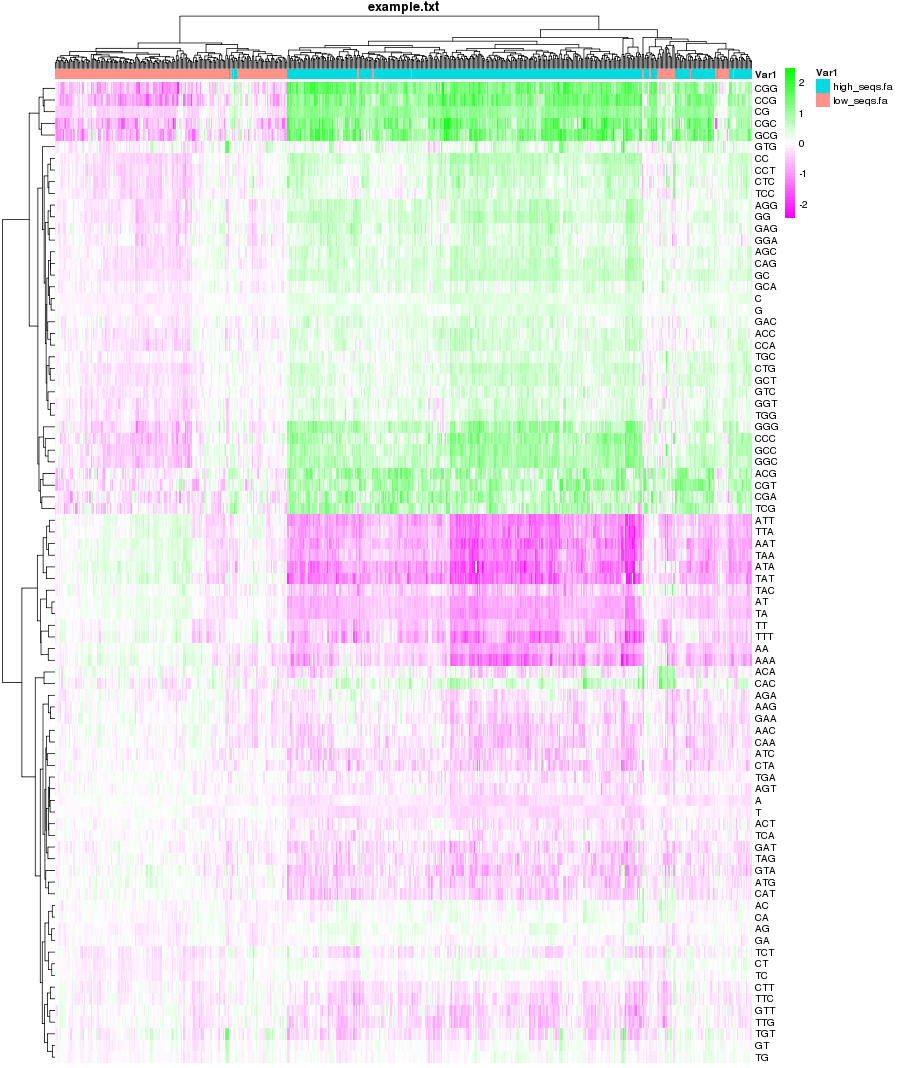
Changelog
See the Github project page commits and releases pages to see the progress of the project.
