Some high throughput sequencers generate sequence fragments of uniform length, but others can contain reads of wildly varying lengths. Even within uniform length libraries some pipelines will trim sequences to remove poor quality base calls from the end.
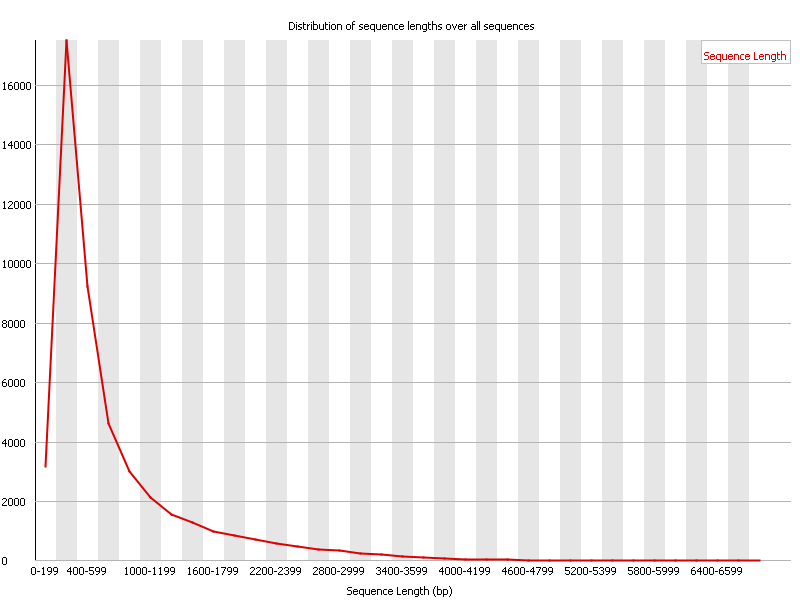
This module generates a graph showing the distribution of fragment sizes in the file which was analysed.

In many cases this will produce a simple graph showing a peak only at one size, but for variable length FastQ files this will show the relative amounts of each different size of sequence fragment.
This module will raise a warning if all sequences are not the same length.
This module will raise an error if any of the sequences have zero length.
For some sequencing platforms it is entirely normal to have different read lengths so warnings here can be ignored.